Inter-Session Alignment and Projection Calculation
After motion correction, CaliAli calculates session projections, aligns sessions, and concatenates them for extraction.
Run:
Output File
This step creates one *_Aligned.mat file plus per-session *_det.mat files. For naming and save-location details, see FAQ output naming.
Evaluating Alignment Performance
While running CaliAli_align_sessions(), check the reported:
- Blood-vessel similarity score
- Spatial correlation after alignment
Quick post-alignment check:
fprintf('BV Score: %.4f\n', CaliAli_options.inter_session_alignment.BV_score);
Alignment_metrics = CaliAli_options.inter_session_alignment.alignment_metrics
plot_alignment_scores(CaliAli_options)
P = CaliAli_options.inter_session_alignment.P;
frame = plot_P(P);
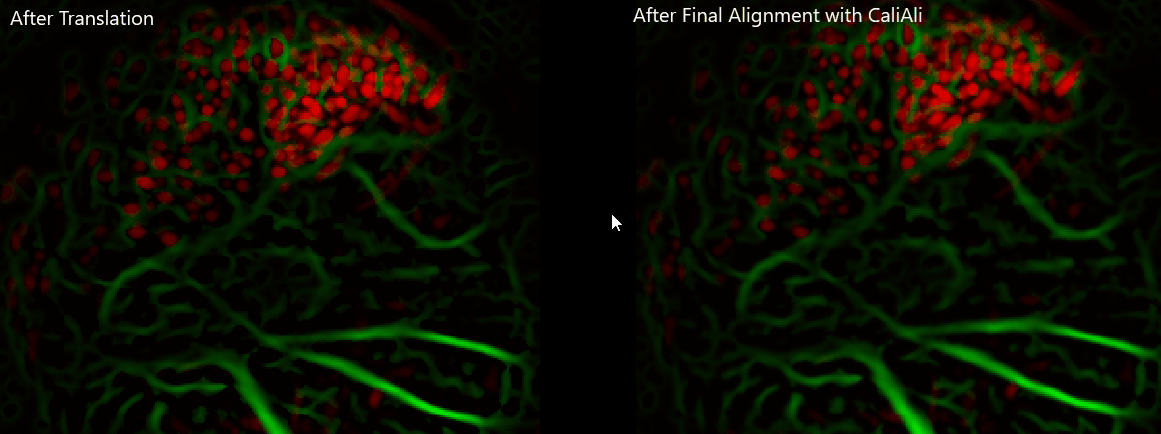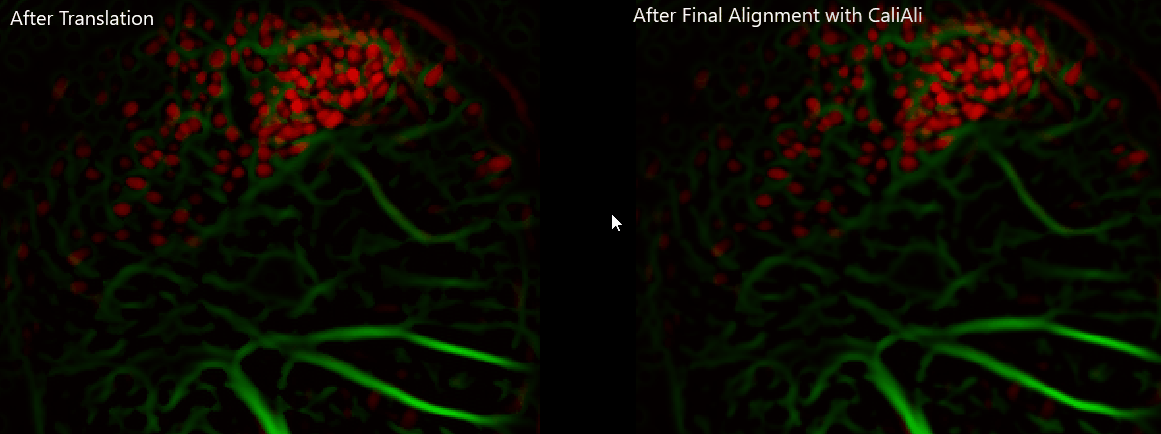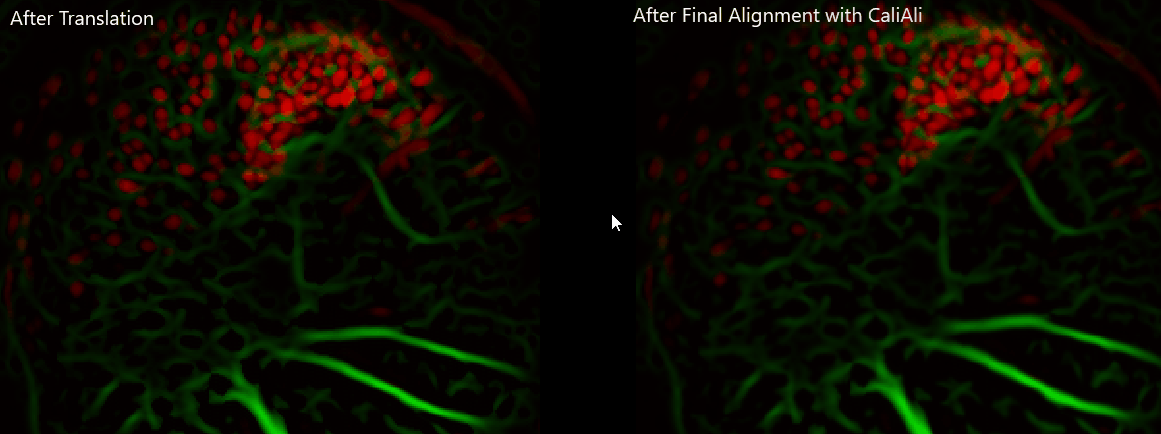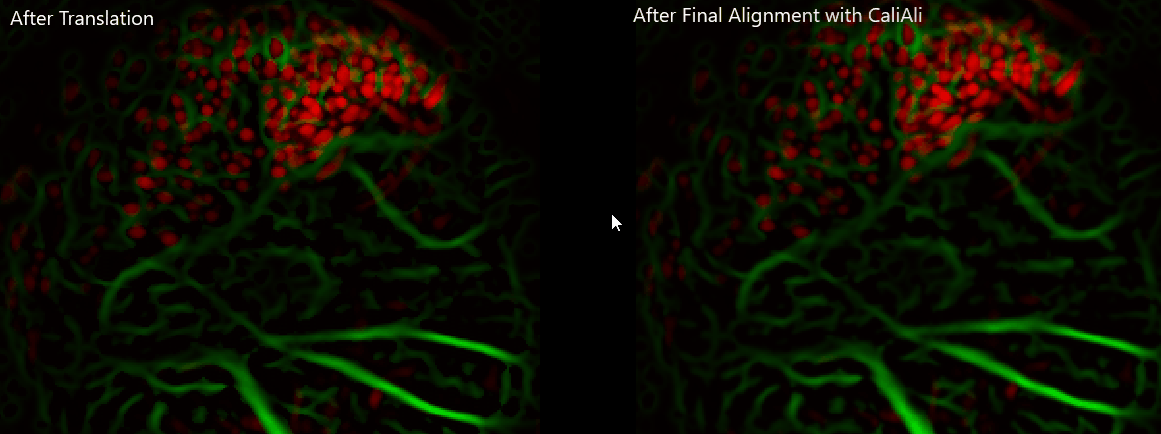
Important
Always visually verify alignment quality before CNMF-E. If sessions remain visibly displaced, downstream extraction quality will be affected.
For advanced details about alignment internals and stored fields, see CaliAli_align_sessions().
After alignment (or single-file projection processing), proceed to Calcium Signal Extraction